2023 | 2022 | 2021 | 2020 | 2019 | 2018 | 2017 | 2016 | 2015 | 2014 | 2013 | 2012 | 2010 | 2009 | 2008 | 2007 | 2006
(click image for PDF, or title for journal link)
2023
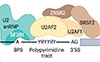
Bradley RK†, Anczuków O†
Nature Reviews Cancer https://doi.org/10.1038/s41568-022-00541-7 (2023) †co-corresponding authors
2022
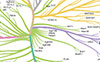
Wang E*,†, Pineda JMB*, Kim WJ*, Chen S, Bourcier J, Stahl M, Hogg SJ, Bewersdorf JP, Han C, Singer ME, Cui D, Erickson CE, Tittley SM, Penson AV, Knorr K, Stanley RF, Rahman J, Krishnamoorthy G, Fagin JA, Creger E, McMillan E, Mak C-C, Jarvis M, Bossard C, Beaupre DM, Bradley RK†, Abdel-Wahab O†
Cancer Cell 41:1-17 (2022) *co-first authors; †co-corresponding authors
Featured in Cancer Discovery.
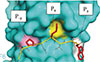
Choudhary GS, Pellagatti A, Agianian B, Smith MA, Bhagat TD, Gordon-Mitchell S, Sahu S, Pandey S, Shah N, Aluri S, Aggarwal R, Aminov S, Schwartz L, Steeples V, Booher RN, Ramachandra M, Samson M, Carbajal M, Pradhan K, Bowman TV, Pillai MM, Will B, Wickrema A, Shastri A, Bradley RK, Martell RE, Steidl RG, Gavathiotis E, Boultwood J, Starczynowski DT, Verma A
eLife 11:e78136 (2022)

North K*, Benbarche S*, Liu B, Pangallo J, Chen S, Stahl M, Bewersdorf JP, Stanley RF, Erickson C, Cho H, Pineda JMB, Thomas JD, Polaski JT, Belleville AE, Gabel AM, Udy DB, Humbert O, Kiem H-P, Abdel-Wahab O†, Bradley RK†
Nature Biotechnology 40:1103–1113 (2022) *co-first authors; †co-corresponding authors
Featured in Nature Reviews Cancer, Cancer Discovery, Nature Biotechnology, and Fred Hutch News.
2021

Udy DB, Bradley RK
Life Science Alliance doi:10.26508/lsa.202101217 (2021)

Clough CA, Pangallo J, Sarchi M, Ilagan JO, North K, Bergantinos R, Stolla MC, Naru J, Nugent P, Kim E, Stirewalt DL, Subramaniam AR, Abdel-Wahab O, Abkowitz JL, Bradley RK†, Doulatov S†
Blood doi:10.1182/blood.2021012652 (2021) †co-corresponding authors

Parrish PCR*, Thomas JD*, Gabel AM, Kamlapurkar S, Bradley RK, Berger AH
Cell Reports 36, 109597 (2021) *co-first authors
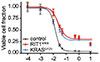

Vichas A*, AK Riley*, Nkinsi NT, Kamlapurkar S, Parrish PCR, Lo A, Duke F, Chen J, Fung I, Watson J, Rees M, Gabel AM, Thomas JD, Bradley RK, Lee JK, Hatch EM, Baine MK, Rekhtman N, Ladanyi M, Piccioni F, Berger AH
Nature Communications 12:4789 (2021) *co-first authors

Beauchamp EM, Leventhal M, Bernard E, Hoppe ER, Todisco G, Creignou M, Gallì A, Castellano CA, McConkey M, Tarun A, Wong W, Schenone M, Stanclift C, Tanenbaum B, Malolepsza E, Nilsson B, Bick AG, Weinstock JS, Miller M, Niroula A, Dunford A, Taylor-Weiner A, Wood T, Barbera A, Anand S; Psaty BM, Desai P, Cho MH, Johnson AD, Loos R; MacArthur DG, Lek M; Neuberg DS, Lage K, Carr SA, Hellstrom-Lindberg E, Malcovati L, Papaemmanuil E, Stewart C, Getz G, Bradley RK, Jaiswal S, Ebert BL
Blood Cancer Discovery 2:500-517 (2021)

Lu SX*, De Neef E*, Thomas JD*, Sabio E, Rousseau B, Gigoux M, Knorr DA, Greenbaum B, Elhanati Y, Hogg SJ, Chow A, Ghosh A, Xie A, Zamarin D, Cui D, Erickson C, Singer M, Cho H, Wang E, Lu B, Durham BH, Shah H, Chowell D, Gabel AM, Shen Y, Liu J, Jin J, Rhodes MC, Taylor RE, Molina H, Wolchok JD, Merghoub T, Diaz LA, Abdel-Wahab O†, Bradley RK†
Cell 184:4032-4047 (2021) *co-first authors; †co-corresponding authors
Featured in The New England Journal of Medicine, Nature, Cancer Discovery, Signal Transduction and Targeted Therapy, GeekWire, Fred Hutch News, and Memorial Sloan Kettering News.
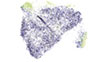
Li S, Simonia Y, Zhuanga S, Gabel A, Mag S, Cheeg J, Islasa L, Cessnaa A, Creaneyg J, Bradley RK, Redwood A, Robinson BW, Newell EW
PNAS 118:e2025570118 (2021)
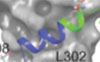
Inoue D*, Polaski JT*, Taylor J*, Castel P, Chen S, Kobayashi S, Hogg SJ, Hayashi Y, Pineda JMB, El Marabti E, Erickson C, Knorr K, Fukumoto M, Yamazaki H, Tanaka A, Fukui C, Lu SX, Durham BH, Liu B, Wang E, Mehta S, Zakheim D, Garippa R, Penson A, Chew G-L, McCormick F, Bradley RK†, Abdel-Wahab O†
Nature Genetics 53:707-718 (2021) *co-first authors; †co-corresponding authors
Featured in Cancer Discovery.
Takao S, Forbes L, Uni M, Cheng S, Pineda JMB, Tarumoto Y, Cifani P, Minuesa G, Chen C, Kharas MG, Bradley RK, Vakoc C, Koche RP, Kentsis A
eLife 10:e65905 (2021)

Chew G-L, Bleakley M, Bradley RK, Malik HS, Henikoff S, Molaro A, Sarthy J
Nature Communications 12:490 (2021)
Featured in Nature Reviews Genetics.

Polaski JT, Udy DB, Escobar-Hoyos LF, Askan G, Leach SD, Ventura A, Kannan R†, Bradley RK†
eLife 10:e62209 (2021) †co-corresponding authors
2020

Taylor J*, Mi X*, North KD*, Binder M*, Penson A, Lasho TL, Knorr K, Haddadin M, Liu B, Pangallo J, Benbarche S, Wiseman DH, Tefferi A, Halene S, Liang Y, Patnaik MM, Bradley RK, Abdel-Wahab O
Blood doi:10.1182/blood.2020006868 (2020) *equal contribution
Featured in Blood.

Escobar-Hoyos LF, Penson A, Kannan R, Cho H, Pan C-H, Singh RK, Apken LH, Hobbs GA, Luo R, Lecomte N, Babu S, Pan FC, Alonso-Curbelo D, Morris JP, Askan G, Grbovic-Huezo O, Ogrodowski P, Bermeo J, Saglimbeni J, Cruz CD, Ho Y-J, Lawrence SA, Melchor JP, Goda GA, Bai K, Pastore A, Hogg SJ, Raghavan S, Bailey P, Chang DK, Biankin A, Shroyer KR, Wolpin BM, Aguirre AJ, Ventura A, Taylor B, Der CJ, Dominguez D, Kümmel D, Oeckinghaus A, Lowe SW, Bradley RK†, Abdel-Wahab O†, Leach SD†
Cancer Cell 38:1–14 (2020) †co-senior authors
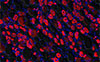
Jones TI, Chew G-L, Barraza-Flores P, Schreier S, Ramirez M, Wuebbles RD, Burkin DJ, Bradley RK, Jones PL
Skeletal Muscle 10:8 (2020)

Rahman MA, Lin K-T, Bradley RK, Abdel-Wahab O, Krainer AR
Genes & Development 34:1–15 (2020)
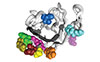
Pangallo J, Kiladjian J-J, Cassinat B, Renneville A, Taylor J, Polaski JT, North K, Abdel-Wahab O, Bradley RK
Blood doi:10.1182/blood.2019002894 (2020)
Featured in Blood.

Thomas JD, Polaski JT*, Feng Q*, De Neef EJ, Hoppe ER, McSharry MV, Pangallo J, Gabel AM, Belleville AE, Watson J, Nkinsi NT, Berger AH, Bradley RK
Nature Genetics 52:84–94 (2020) *equal contribution
Featured in Yahoo! News, FierceBiotech, GeekWire, and Fred Hutch News.
2019

Inoue D*, Chew G-L*, Liu B, Michel BC, Pangallo J, D’Avino AR, Hitchman T, North K, Lee SC-W, Bitner L, Block A, Moore AR, Yoshimi A, Escobar-Hoyos L, Cho H, Penson A, Lu SX, Taylor J, Chen Y, Kadoch C, Abdel-Wahab O†, Bradley RK†
Nature 574:432–436 (2019) *co-first authors; †co-corresponding authors
Yoshimi A, Lin K-T, Wiseman DH, Rahman MA, Pastore A, Wang B, Lee SC-W, Micol J-B, Zhang XJ, de Botton S, Penard-Lacronique V, Stein EM, Cho H, Miles RE, Inoue D, Albrecht TR, Somervaille TCP, Batta K, Amaral F, Simeoni F, Wilks DP, Cargo C, Intlekofer AM, Levine RL, Dvinge H, Bradley RK, Wagner EJ, Krainer AR, Abdel-Wahab O
Nature 574:273–277 (2019)

Dvinge H†, Guenthoer J, Porter PL, Bradley RK†
Genome Research 29:1591–1604 (2019) †co-corresponding authors

Chew G-L*, Campbell A*, De Neef E, Sutliff NA, Shadle SC, Tapscott SJ†, Bradley RK†
Developmental Cell 50:658–671.e7 (2019) *co-first authors; †co-corresponding authors
Featured in Nature Reviews Cancer, FierceBiotech, GeekWire, and Fred Hutch News.
Palangat M, Anastasakis DG, Fei DL, Lindblad KE, Bradley RK, Hourigan CS, Hafner M, Larson DR
Genes & Development 33:1–16 (2019)
Jagannathan S†, Ogata Y, Gafken PR, Tapscott SJ†, Bradley RK†
eLife 8:e41740 (2019) †co-corresponding authors
2018
Blazquez L, Emmett W, Faraway R, Pineda JMB, Bajew S, Gohr A, Haberman N, Sibley CR, Bradley RK, Irimia M, Ule J
Molecular Cell 72:496–509 (2018)
Fei DL, Zhen T, Durham B, Ferrarone J, Zhang T, Garrett L, Yoshimi A, Abdel-Wahab O, Bradley RK, Liu P, and Varmus H
PNAS doi:10.1073/pnas.1812669115 (2018)

Lee SC*, North K*, Kim E*, Jang E, Obeng E, Lu SX, Liu B, Inoue D, Yoshimi A, Ki M, Yeo M, Zhang XJ, Kim MK, Cho H, Chung YR, Taylor J, Durham BH, Kim YJ, Pastore A, Monette S, Palacino J, Seiler M, Buonamici S, Smith PG, Ebert BL, Bradley RK†, Abdel-Wahab O†
Cancer Cell 34:225–241 (2018) *co-first authors; †co-corresponding authors
Featured in Cancer Discovery.

Chang C-J, Kotini AG, Olszewska M, Georgomanoli M, Teruya-Feldstein J, Sperber H, Sanchez R, DeVita R, Martins TJ, Abdel-Wahab O, Bradley RK, Papapetrou EP
Stem Cell Reports 10:1610–1624 (2018)
Pineda JMB, Bradley RK
Genes & Development 32:577–591 (2018)
2017

Jagannathan S, Bradley RK
Genes & Development 31:1067–1068 (2017)
Feng Q, Jagannathan S, Bradley RK
Molecular Cell 67:239–251 (2017)
Featured in Bioessays.
2016

Fei DL†, Motowski H, Chatrikhi R, Prasad S, Yu J, Gao S, Kielkopf CL†, Bradley RK†, Varmus H†
PLoS Genetics 12(10):e1006384 (2016) †co-corresponding authors
Uo T, Dvinge H, Sprenger CC, Bradley RK, Nelson PS, Plymate SR
Oncogene 36:1440–1450 (2016)

Jagannathan S, Bradley RK
Genome Research 26:1639–1650 (2016)

Jagannathan S*, Shadle S*, Resnick R, Snider L, Tawil RN, van der Maarel SM, Bradley RK†, Tapscott SJ†
Human Molecular Genetics 25:4419–4431 (2016) *co-first authors; †co-senior authors

Dvinge H*, Kim E*, Abdel-Wahab O†, Bradley RK†
Nature Reviews Cancer 16:413–430 (2016) *co-first authors; †co-senior authors
Lee SC*, Dvinge H*, Kim E, Cho H, Micol JB, Chung YR, Durham BH, Yoshimi A, Kim YJ, Thomas M, Lobry C, Chen CW, Pastore A, Taylor J, Wang X, Krivtsov A, Armstrong SA, Palacino J, Buonamici S, Smith PG, Bradley RK†, Abdel-Wahab O†
Nature Medicine 22:672–678 (2016) *co-first authors; †co-senior authors
Featured in F1000 and Cancer Discovery.
Inoue D, Bradley RK, Abdel-Wahab O
Genes & Development 30:989–1001 (2016)
2015

The Cancer Genome Atlas Research Network
Cell 163:1011–1025 (2015)
Hickey TE, Irvine CM, Dvinge H, Tarulli GA, Hanson AR, Ryan NK, Pickering MA, Birrell SN, Hu DG, Mackenzie PI, Russell R, Caldas C, Raj GV, Dehm SM, Plymate SR, Bradley RK, Tilley WD, Selth LA
Oncotarget 10.18632/oncotarget.6296 (2015)

Robinson D, Van Allen EM, et al
Cell 161:1215–1228 (2015)
Kim E*, Ilagan JO*, Liang Y*, Daubner GM*, Lee SC-W, Ramakrishnan A, Li Y, Chung YR, Micol J-B, Murphy M, Cho H, Kim M-K, Zebari AS, Aumann S, Park CY, Buonamici S, Smith PG, Deeg HJ, Lobry C, Aifantis I, Modis Y, Allain FH-T, Halene S†, Bradley RK†, Abdel-Wahab O†
Cancer Cell 27:617–630 (2015) *co-first authors; †co-senior authors
Featured in Nature Reviews Cancer, Cancer Cell, and Fred Hutch News.
Dvinge H, Bradley RK
Genome Medicine 7:45 (2015)
Feng Q, Snider L, Jagannathan S, Tawil R, van der Maarel SM, Tapscott SJ†, Bradley RK†
eLife 4:e04996 (2015) †co-senior authors
Featured in F1000.
2014
Dvinge H, Ries RE, Ilagan JO, Stirewalt DL, Meshinchi S, Bradley RK
PNAS 111:16802–16807 (2014)
Featured in F1000 and Fred Hutch News.
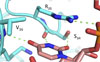
Ilagan JO*, Ramakrishnan A*, Hayes B, Murphy ME, Zebari AS, Bradley P, Bradley RK
Genome Research doi:10.1101/gr.181016.114 (2014) *co-first authors
2013

Hubert CG*, Bradley RK*, Ding Y, Toledo CM, Herman J, Skutt-Kakaria K, Girard EJ, Davison J, Berndt J, Corrin P, Hardcastle J, Basom R, Delrow JJ, Webb T, Pollard SM, Lee J, Olson JM, Paddison PJ
Genes & Development 27(9):1032–1045 (2013) *co-first authors
Featured in Cancer Discovery.

Xi Z, Yuguo W, Bradley RK, Sugumaran M, Marx CJ, Rest JS, Davis CC
PLoS Genetics 9(2):e1003265 (2013)
Klattenhoff C, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML, Ding H, Butty VL, Torrey L, Haas S, Abo R, Tabebordbar M, Lee RT, Burge CB, Boyer LA
Cell 152(3):570–583 (2013)
2012

Xi Z, Bradley RK, Wurdack KJ, Wong KM, Sugumaran M, Bomblies K, Rest JS, Davis CC
BMC Genomics 13:227 (2012)
Featured in The Economist and Scientific American.
Satija R, Bradley RK
Genome Research 22:656–665 (2012)

Bradley RK, Merkin JM, Lambert N, Burge CB
PLoS Biology 10:e1001229 (2012)
Featured in Molecular Cell and Genome Biology.
2010

Bradley RK*, Li XY*, Trapnell C, Davidson S, Pachter L, Chu HC, Tonkin LA, Biggin MD, Eisen MB
PLoS Biology 8:e1000343 (2010) *co-first authors
Featured in PLoS Biology.
2009
Bradley RK, Holmes I
PLoS Computational Biology 5(8):e1000483 (2009)

Bradley RK, Uzilov AV, Skinner ME, Bendana YR, Barquist L, Holmes I
PLoS One 4(8):e6478 (2009)

Bradley RK, Roberts A, Smoot M, Juvekar S, Do J, Dewey C, Holmes I, Pachter L
PLoS Computational Biology 5:e1000392 (2009)
2008

Varadarajan A, Bradley RK, Holmes I
Genome Biology 9:R147 (2008)

Bradley RK, Pachter L, Holmes I
Bioinformatics 24:2677–2683 (2008)
2007
Drosophila 12 Genomes Consortium
Nature 450:203–218 (2007)

Bradley RK, Holmes I
Bioinformatics 23:3258–3262 (2007)
2006

Klosterman PS, Uzilov AV, Bendana YR, Bradley RK, Chao S, Kosiol C, Goldman N, Holmes I
BMC Bioinformatics 7:428 (2006)